Home
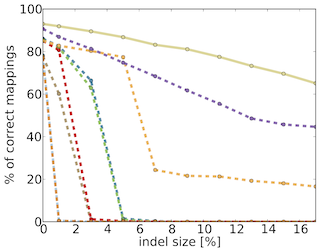
TreQ is a read mapper for high-throughput DNA sequencing reads, in particular one to several hundred nucleotides in length, and for large edit distance between sequencing read and match in the reference genome. In contrast to existing read mappers, TreQ can cope particularly well with indels, as demonstrated in the figure below for single-best hit recall of 200nt reads simulated from the human reference genome. TreQ (solid line) performs best at a running time comparable to BWA at large edit distance settings. SSAHA2 (purple) shows reasonable accuracy, but is five times slower. This makes TreQ an excellent choice for analyzing genetic variants in low-coverage situations and without the need for paired-end sequencing. TreQ will be released under the GPL upon publication.
TreQ is developed and maintained at Rutgers, The State University of New Jersey, Department of Computer Science and the BioMaPS Institute for Quantitative Biology.



