MSRE-HTPrimer_Manual
**The user manual of MSRE-HTPrimer can be downloaded in PDF format from here... (sourceforge.net)
**
TABLE OF CONTENTS
1. What is MSRE-HTPrimer?
MSRE-HTPrimer is a high-throughput and genome wide primer design pipeline for epigenetic and genomic sequencing primer design for hundreds to thousands of target sequences in a single run with great accuracy, sensitivity and success rate. It offers easy to handle input and output options, which is especially designed for non-expert users.
MSRE-HTPrimer provides significant improvements over existing solutions with following unique features 1) parallel primer design for several target sequences, 2) primer selection and ranking based on custom filtering criteria, 3) links to Insilico-PCR for each resulting primer pair, 4) visualization of the primer design result in UCSC genome browser, and 5) takes SNPs and repeats into consideration for primer design and automatically discard primer pairs which falls in repeat region or contain SNPs. The pipeline is equipped with an intuitive web interface and multiprocessing capability and provides custom inputs and parameters to design target specific primers. MSRE-HTPrimer is a user-friendly and standalone tool, which is available within a fully configured Virtual machine. It does not require any installation or configuration except VirtualBox (http://www.virtualbox.com).
2. System requirements
Virtual box from https://www.virtualbox.org/
Operating system
Linux
Mac OSX 10.6 or later
Windows PC
3. How to obtain MSRE-HTPrimer?
MSRE-HTPrimer is a web-based standalone tool available within a fully configured Virtual box and can be downloaded from https://sourceforge.net/p/msrehtprimer/wiki/Virtual_Machine and can be easily run on any operating system. With Virtual machine no installation and configuration required.
Once virtual machine of MSRE-HTPrimer is obtained then follow these steps to run the MSRE-HTPrimer tool.
Step1.1: Download and install the Virtual Box (version 5.1.2) from http://www.oracle.com/technetwork/server-storage/virtualbox/downloads/index.html#vbox
Step1.2: After installation of Virtual Box download and install the Virtual Box Extension Pack (version 5.1.2) from http://download.virtualbox.org/virtualbox/5.1.2/Oracle_VM_VirtualBox_Extension_Pack-5.1.2.vbox-extpack
Step2: Import MSRE-HTPrimer Virtual Machine file into Virtual Box
Step3: Login into MSRE-HTPrimer Virtual machine with username = testuser and password = testuser
Step4: Open Firefox or any other web browser and open the query page of MSRE-HTPrimer with http://localhost/msre-htprimer
Step5: Run the MSRE-HTPrimer with the test data sets.
Step6: For new data analysis with MSRE-HTPrimer, prepare the Target file and run the primer design.
4. MSRE-HTPrimer web interface description
4.1 MSRE-HTPrimer query page
User can run primer design with MSRE-HTPrimer by providing input options and files from query page as shown in Figure 1.
Following input parameters and input files are required.
Input 1:
Genome information parameters are required to download genome fasta sequence and annotation files (RefSeq gene, common SNPs, CpG island and known repeat elements) from UCSC genome browser (http://genome.ucsc.edu/index.html)
1) Select genome name from first drop down menu (Human or Mouse). The default genome is Human.
2) Select genome assembly version from second drop down menu (default genome assembly is hg38)
3) Select the dbSNP build to download the corresponding common SNPs from UCSC genome browser. Default is 142 for Human, genome assembly hg19.
**Input 2: **
Upload a target file in BED format. One file for each target region. This file consists of 4 columns 1) chromosome, 2) start position, 3) end position and 4) target ID. User can give any number of target region in a single run.
Input 3:
Third input is the Primer3 input parameters for primer design. This file can be modified as per the requirement and upload. This is an optional input, if user does not provide then MSRE-HTPrimer uses the default setting of Primer3.
Input 4:
This input is only required fo MSRE-PCR primer design, user can either enter type-II enzyme is the input box (one enzyme per line) or alternatively can upload a text file which contains one enzyme per line.
Input 5:
To provide flexibility in primer design process, MSRE-HTPrimer provides some useful input options for optimized and speific primer design and selection. Under these parameters user can define
- Maximum primer pairs to return for each target region. Default 10
- Product size: Minimum, Optimum and Maximum. Default 150, 250 and 320 respectively.
- Primer annealing temperature. Default 52, 60 and 65 for Minimum, Optimum and Maximum temperature respectively.
- Primer size: Minimum, Optimum and Maximum. Default 22, 28 and 36 bp respectively.
Input 6:
This is unique feature of MSRE-HTPrimer to provide primer selection quality matrix for final primer pair selection, which helps to reduce the post selection process. User can define various filtering criteria for each output column of the MSRE-HTPrimer and then tool automatically selection primers from the whole output. This is optional input.

Figure 1: shows MSRE-HTPrimer query page to define all parameters and upload input files for primer design
4.2 MSRE-HTPrimer result page
MSRE-HTPrimer results all primer pairs in a summary table, which is available in HTML to display (as shown in Figure 2) and in TEXT and HTML format to download. Moreover, MSRE-HTPrimer has seamlessly integrated the UCSC genome browser and UCSC In-Silico PCR visualization in the result page. All resulting primer pairs of a target region is visualized in UCSC genome browser as shown in Figure 3. For each primer pair In-Silico PCR is displayed within the result page of MSRE-HTPrimer as shown Figure 4.

Figure 2: shows MSRE-HTPrimer primer pair output summary table (http://localhost/msre-htprimer).
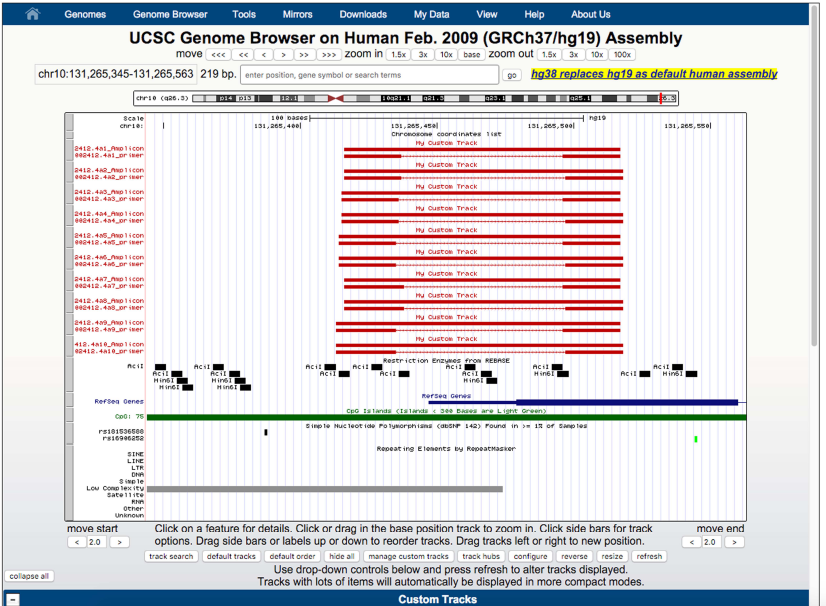
Figure 3: shows the visualization of primer pairs in the UCSC genome browser within the MSRE-HTPrimer result page Go to UCSC genome browser
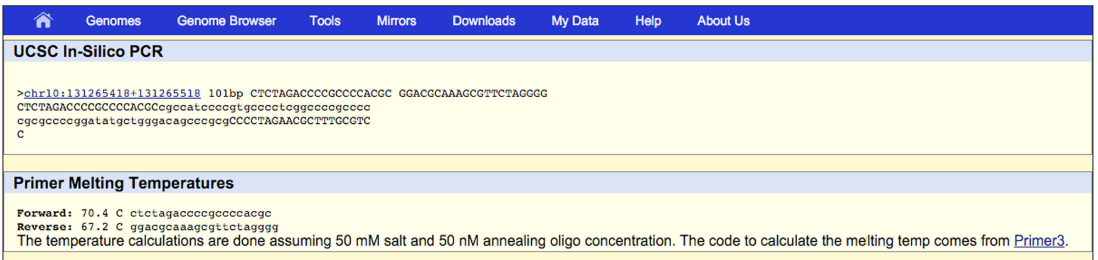
Figure 4: shows the visualization of primer pairs in the UCSC In-Silico PCR within the MSRE-HTPrimer result page Go to UCSC In-Silico PCR
4.3 Run MSRE-HTPrimer with test input
To validate the installation of the MSRE-HTPrimer pipeline it can be run with a small test data set. The test data set for MSRE-PCR and Genomic-PCR can be obtained from https://sourceforge.net/projects/msrehtprimer/files/test_data.zip
and run the following command to uncompress the file:
unzip test_data.zip
Note that after uncompressing the .zip file, a new folder will be created named test_data. Now upload these files on the MSRE-HTPrimer query page (http://localhost/msre-htprimer) and run the primer design.
5. MSRE-HTPrimer inputs description
MSRE-HTPrimer requires four input files:
5.1 Target BED file
This file contains the genomic coordinates for all target sequences (one line for each target sequence). It consists of four tab-delimited columns: 1) chromosome, 2) start coordinate, 2) end coordinate and 4) a unique ID for each target region as shown in Table below.
chr2 241454334 241457334 Target1
chr3 10155818 10158818 Target2
chr5 118813546 118816546 Target3
chr5 148183848 148186848 Target4
chr5 112098954 112101954 Target5
chr15 89059082 89062082 Target6
chr19 1154297 1157297 Target7
chrY 25386895 25389895 Target8
5.2 Primer3 parameter file
This text file contains the parameters and values for the Primer3 tool. It is optional and if not provided, MSRE-HTPrimer will use default Primer3 parameters as shown below:
PRIMER_TASK=generic
PRIMER_MISPRIMING_LIBRARY=
PRIMER_MIN_TM=65.0
PRIMER_OPT_TM=70.0
PRIMER_MAX_TM=75.0
PRIMER_MIN_GC=20.0
PRIMER_MAX_GC=100.0
PRIMER_NUM_RETURN=5000
PRIMER_MIN_SIZE=16
PRIMER_OPT_SIZE=21
PRIMER_MAX_SIZE=30
PRIMER_PRODUCT_SIZE_RANGE=50-150
SEQUENCE_ID=TS001
SEQUENCE_TEMPLATE=
PRIMER_PICK_LEFT_PRIMER=1
PRIMER_PICK_RIGHT_PRIMER=1
PRIMER_PICK_INTERNAL_OLIGO=1
PRIMER_PICK_ANYWAY=1
PRIMER_THERMODYNAMIC_OLIGO_ALIGNMENT=0
PRIMER_THERMODYNAMIC_TEMPLATE_ALIGNMENT=0
5.3 Restriction enzyme file
This input file is only required for MSRE-PCR primer design. Each line contains an enzyme name as per nomenclature and multiple enzymes are allowed in a single run as shown below:
HpaII
Hin6I
AciI
HpyCH4IV
5.4 Custom primer selection quality matrix
MSRE-HTPrimer supports further selection of primer pairs based on user defined selection criteria. A custom quality-filtering matrix can be provided as input file. As shown in Table below, the user can define a set of selection criteria and rank them using a scale of 1-10. MSRE-HTPrimer assigns these ranks to the primer pairs for all target sequences. If this input is not provided then primer pairs are returned based on Primer3 ranking. MSRE-HTPrimer supports mathematical operators, including “>”, “<”, “>=”, “<=” and “-“. Any column header of the MSRE-HTPrimer output file can be used as parameter. The primer quality level represents the rank associated with each of the output parameters in its respective row.
Table: Custom quality filter matrix with ten quality levels ranking the designed primer independent of the primer3 level, but dependent on amplicon size, amount of cutsites and gene distance.

All headers consist of two major parts, origin (Fp=forward primer, Lp=left primer, Rp=reverse primer/right primer, Amp=amplicon, Hyb=hybridization oligo) and short description. Primer_Quality_Level=user defined rank; Tm=melting temperature of origin; Gc_%=GC percentage in DNA sequence of origin; Any_Compl=stability of any basepairing of origin to itself; 3'_Compl= stability of any basepairing of the 3' end of the origin to itself; Size=size of origin in basepairs (Bp); Repeat_In_Bp=allowed Bp of repeats in origin; Snp_Pos_From_3'=distance of closest SNP position to 3’end inside the origin sequence in basepairs; Amp_Sum_Cutsites_Primer=amount of cutsites in FP and RP; Amp_Sum_Cutsites_Between_Primers=amount of cutsites in amplicon except for FP and RP.
6. How to use MSRE-HTPrimer?
To use MSRE-HTPrimer for primer design user
Open web browser and type the following url into browser (http://localhost/msre-htprimer) and upload required inputs files, change default parameters if required and run the primer design.
7. Contact Information
Ram Vinay Pandey
ramvinay.pandey@gmail.com


