TCellXTalk is a comprehensive database of experimentally detected phosphorylation, ubiquitination and acetylation sites in human T cells.
The web-app at www.TCellXTalk.org makes TCellXTalk accessible from Internet, and enables the in silico prediction of potential co-modified peptides to facilitate their experimental detection, using targeted or directed mass spectrometry, for the study of protein post-translational modification cross-talk.
More detailed information on TCellXTalk and the people at the CSIC/UAB Proteomics Laboratory behind it can be obtained at https://www.tcellxtalk.org/about/
- Casanovas, A., Gallardo, Ó., Carrascal, M., Abian, J., TCellXTalk facilitates the detection of co-modified peptides for the study of protein post-translational modification cross-talk in T cells. Bioinformatics 2019, 35-8, 1404–1413, DOI 10.1093/bioinformatics/bty805 ( https://doi.org/10.1093/bioinformatics/bty805 )
Features
- On-line at https://www.TCellXTalk.org
- PTMs Information
- Phosphorylation, Ubiquitination and Acetylation sites in human T cells
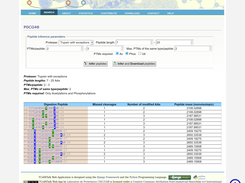
- PTM CorssTalk information and prediction
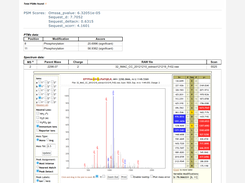
- Mass Spectra
- Proteomics Data
- Python-Django Web-App
- MySQL Database
- README.md file: https://sourceforge.net/p/lp-csic-uab/tcellxtalk/code/ci/default/tree/README.md
- Guide to Devs: https://kutt.it/TCellXTalkDevGuide